| Japanese top page / English top page |
PAICS
Parallelized ab initio Calculation System based on FMO
|
|
|||
|
| |||
| In this page, we show the theoretical study with the fragment molecular orbital method for the interaction between prion protein and a small molecule using PPTs. These PPTs were used for the previous presentations. The following results have been pablished [1]. | |||
|
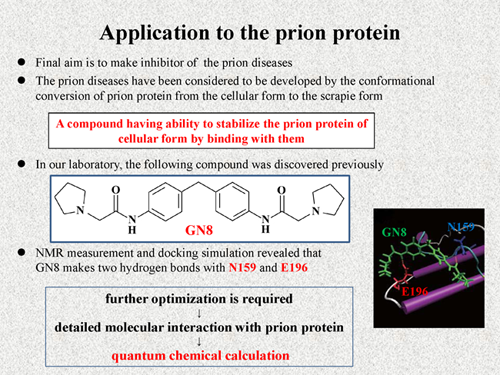
I show the application of the fragment molecular orbital (FMO) method [2] to the prion protein, in which our aim is to make inhibitor of the prion diseases. The prion diseases have been considered to be developed by the conformational conversion from the cellular form to the scrapie form. Thus, we have to find a compound having ability to stabilize the prion protein of cellular form by binding to them.
In our laboratory, the following compound was discovered previously [3]. We call this compound "GN8". The GN8 has ability to inhibit the conformational conversion of prion protein. Our NMR measurement and docking simulation revealed that GN8 makes two hydrogen bonds with N159 and E196, and bridging conformation is formed like this figure. Although the GN8 is a potential curative agent for prion diseases, further optimization is required. For this purpose, it's important to understand the molecular interaction between prion protein and GN8 in more detail. Thus, we employed the quantum chemical calculation using the FMO method. |
|||
 |
|||
|
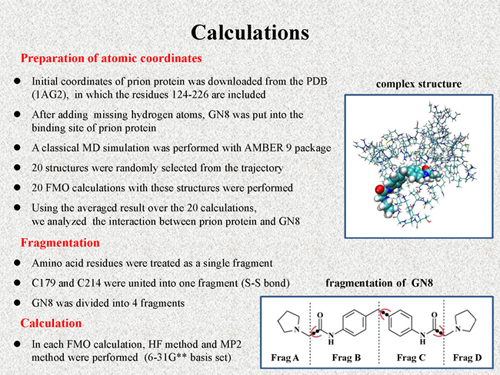
In the case of quantum chemical calculations, atomic coordinates of the system have to be prepared at first. In this work, we prepared them as the following manner. Initial coordinates of prion protein was downloaded from the PDB (1AG2),
in which the residues form 124 to 226 are included. After adding the missing hydrogen atoms, GN8 was put into the binding cite of prion protein. A classical MD simulation was performed with solvent condition using AMBER 9 package, from which 20 structures were randomly selected from the trajectory. FMO calculations with these 20 structures were performed.
Finally, using the average values over the 20 calculations, we analyzed the interaction between prion protein and GN8.
In these FMO calculations, amino acid residues were treated as a single fragment. But, C179 and C214 were treated as one fragment because they are connected with each other by S-S bond. And, GN8 was divided into 4 fragments, as shown in this figure. In each FMO calculation, HF level calculation and MP2 level calculation were performed using 6-31** basis set. |
|||
 |
|||
|
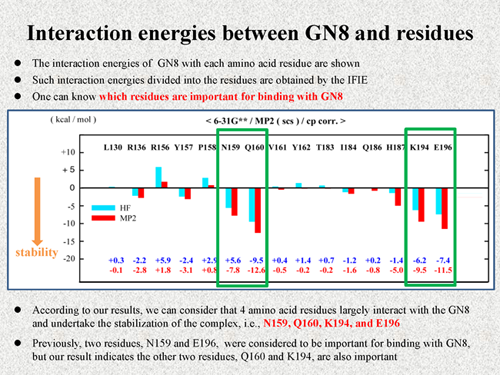
Let's see the results of calculations. First, I show the interaction energies between GN8 and each amino acid residue. Such interaction energies divided into residues are obtained from the inter fragment interaction energy (IFIE) [4]. Using these data, we can know which residues are important for binding with GN8. In this figure, blue sticks are results of the HF calculation and red stick are results of the MP2 calculation, i.e., red results include van der Waals interaction. The lower direction of the interaction energy means undertaking the stabilization of the complex. Here, I show only the results of residues within 3 A from the GN8.
According to our results, we can consider that 4 amino acid residues largely interact with the GN8 and undertake the stabilization of the complex. Previously, two residues of N159 and E196 were considered to be important for the binding with GN8. But, our calculation indicates the other two residues Q160 and K194 are also important. As shown here, using the IFIE, we can know which residues are important for binding with the ligand molecule. |
|||
 |
|||
|
Next, I show another result.
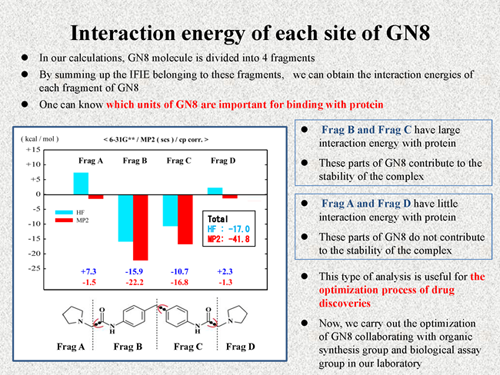
In our calculations, GN8 molecule is divided into 4 fragments. Thus, by summing up the IFIE belonging to these fragments, we can obtain the interaction energies of each fragment of GN8. By this analysis, one can know which units of GN8 are important for binding with prion protein. The results are shown in this figure.
According to our result, Frag B and Frag C have large interaction energy with protein. Thus, we can consider that these parts of GN8 contribute to the stability of the complex. On the other hand, Frag A and Frag D have little interaction energy with protein. We consider that these parts of GN8 do not contribute to the stability of the complex. Like this, with the IFIE, we can know which units of the ligand molecules are important for binding with protein. We expect this type of analysis is useful for the optimization process of drug discoveries. Now, we carry out the optimization of GN8 with organic synthesis group and biological assay group in our laboratory. |
|||
 |
|||
| [1] | Theoretical study of the prion protein based on the fragment molecular orbital method, T. Ishikawa, T. Ishikura, and K. Kuwata, J. Comput. Chem., 30 (2009) 2594-2601 (DOI: 10.1002/jcc.21265) | ||
| [2] | Fragment molecular orbital method: an approximate computational method for large molecules, K. Kitaura, E. Ikeo, T. Asada, T. Nakano, and M. Uebayasi, Chem. Phys. Lett., 313 (1999) 701-706 (DOI: 10.1016/S0009-2614(99)00874-X) | ||
| [3] | Hot spots in prion protein for pathogenic conversion, K. Kuwata, N. Nishida, T. Matsumoto, Y. O. Kamatari, J. Hosokawa-Muto, K. Kodama, H. K. Nakamura, K. Kimura, M. Kawasaki, Y. Takakura, S. Shirabe, J. Takata, Y. Kataoka, and S. Katamine, Proc. Natl. Acad. Sci. USA, 104 (2007) 11921-11926 (DOI: 10.1073/pnas.0702671104) | ||
| [4] | Fragment molecular orbital method: use of approximate electrostatic potential, T. Nakano, T. Kaminuma, T. Sato, K. Fukuzawa, Y. Akiyama, M. Uebayasi, and K. Kitaura, Chem. Phys. Lett., 351 (2002) 475-480 (DOI: 10.1016/S0009-2614(01)01416-6) | ||
|